In this post I want to find out how the Fisherfaces algorithm performs on a classification task like gender classification. There is no specific reason for choosing gender as classification task, so please do not feel offended.
Dataset
I have used the celebrity dataset from my last post and downloaded images for Emma Watson, Naomi Watts and Jennifer Lopez.
So the database has 8 male and 5 female subjects, each with 10 images. All images were chosen to have a frontal face perspective and have been cropped to a size of 140x140 pixels,
just like this set of images.
You can easily find some images yourself using Google Images, then locate the eye positions and use a tool like crop_face.py to geometrically normalize it.
Experiments
You can get the code from github by either cloning the repository:
$ git clone git@github.com:bytefish/facerec.git
or downloading the latest version as a tar or zip archive.
Then startup Octave and make the functions available:
$ cd facerec/m
$ octave
octave> addpath(genpath("."));
octave> fun_fisherface = @(X,y) fisherfaces(X,y); % no parameters needed
octave> fun_predict = @(model,Xtest) fisherfaces_predict(model,Xtest,1); % 1-NN
subject-dependent cross-validation
A Leave-One-Out Cross validation shows that the Fisherfaces method can very accurately predict, wether a given face is male or female:
octave> [X y width height names] = read_images("/home/philipp/facerec/data/gender/");
octave> cv0 = LeaveOneOutCV(X,y,fun_fisherface,fun_predict,1)
129 1 0 0
There are 129 correct predictions and only 1 false prediction, which is a recognition rate of 99%.
subject-independent cross-validation
The results we got were very promising, but they are probably misleading. Have a look at the way a leave-one-out and k-fold cross-validation splits a Dataset D with 3 classes each with 3 observations:
o0 o1 o2 o0 o1 o2 o0 o1 o2
c0 | A B B | c0 | B A B | c0 | B B A |
c1 | A B B | c1 | B A B | c1 | B B A |
c2 | A B B | c2 | B A B | c2 | B B A |
Allthough the folds are not overlapping (training data is never used for testing) the training set contains images of persons we want to know the gender from. So the prediction may depend on the subject and the method finds the closest match to a persons image, but not the gender. What we aim for is a split by class:
o0 o1 o2 o0 o1 o2 o0 o1 o2
c0 | A A A | c0 | B B B | c0 | B B B |
c1 | B B B | c1 | A A A | c1 | B B B |
c2 | B B B | c2 | B B B | c2 | A A A |
With this strategy the cross-validation becomes subject-independent, because images of a subject are never used for learning the model.
We restructure our folder hierarchy, so that it now reads: <group>/<subject>/<image>:
philipp@mango:~/facerec/data/gender$ tree .
.
|-- female
| |-- crop_angelina_jolie
| | |-- crop_angelina_jolie_01.jpg
| | |-- crop_angelina_jolie_02.jpg
| | |-- crop_angelina_jolie_03.jpg
[...]
`-- male
|-- crop_arnold_schwarzenegger
| |-- crop_arnold_schwarzenegger_01.jpg
| |-- crop_arnold_schwarzenegger_02.jpg
| |-- crop_arnold_schwarzenegger_03.jpg
[...]
15 directories, 130 files
The function for reading the dataset needs to return both, the group and the subject of each observation. The cross-validation then predicts on the groups and leaves one subject out in each iteration. read_images.m can be easily extended to read_groups.m and a simple subject-independent cross-validation is quickly written.
On a subject-independent cross-validation the Fisherfaces method is able to predict 128 out of 130 gender correctly, which is a recognition rate of 98%:
octave> [X y g group_names subject_names width height] = read_groups("/home/philipp/facerec/data/gender/");
octave> cv0 = LeaveOneClassOutCV(X,y,g,fun_fisherface,fun_predict,1)
cv0 =
128 2 0 0
Results
Let's have a look at the features identified by the Discriminant Analysis. Since it's a binary classification, there's only one Fisherface. We'll plot it with:
fisherface = fisherfaces(X,g);
fig = figure('visible','off');
imagesc(cvtGray(fisherface.W,width,height));
% remove offsets, set to pixels and scale accordingly
set(gca,'units','normalized','Position',[0 0 1 1]);
set(gcf,'units','pixels','position',[0 0 width height]);
print("-dpng",sprintf("-S%d,%d",width,height),"fisherface.png");
And the Fisherface shows, which image features have been used to classify the samples:
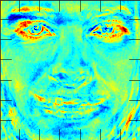
The recognition rate for a subject-independent cross-validation was 98.5%, which is only slightly below the 99.2% for a subject-dependent validation.
Of course, the experimental results should be validated on larger datasets and shouldn't be taken for granted.